spectrum_utils: A Python Package for Mass Spectrometry Data Processing and Visualization | Analytical Chemistry

Assessing production variability in empty and filled adeno-associated viruses by single molecule mass analyses: Molecular Therapy - Methods & Clinical Development
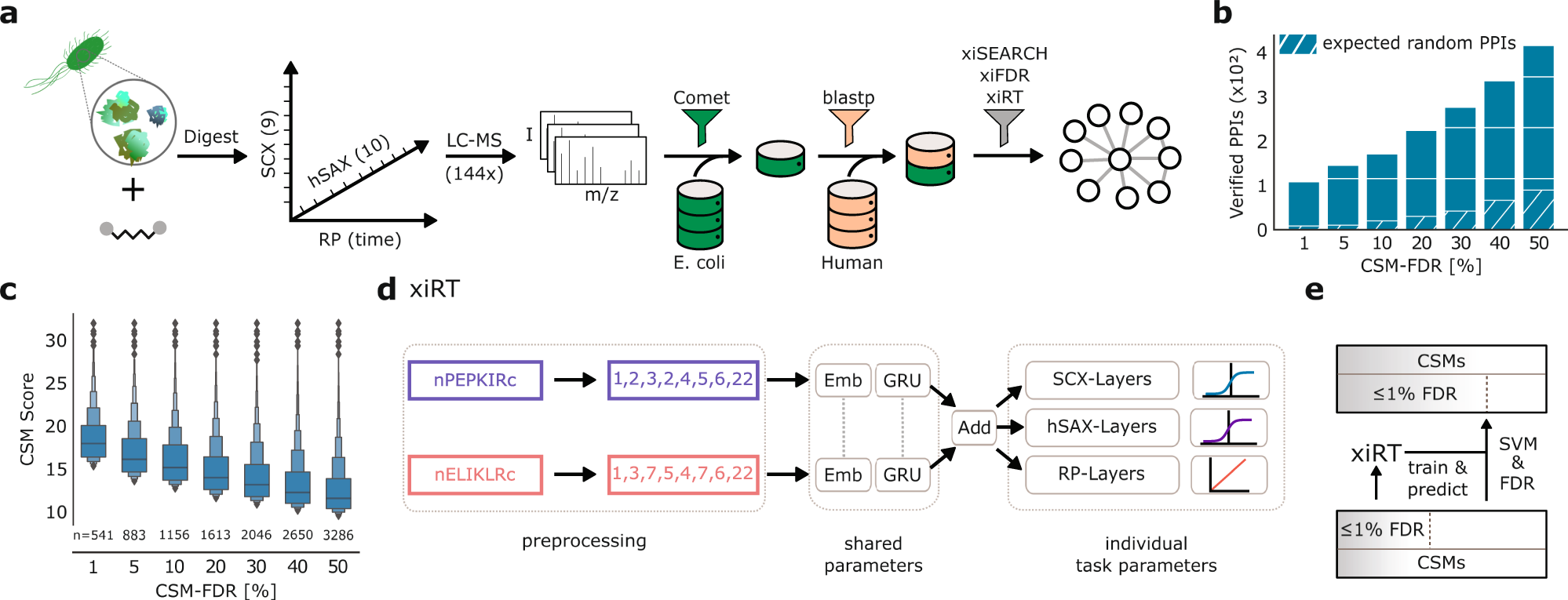
Retention time prediction using neural networks increases identifications in crosslinking mass spectrometry | Nature Communications

Internal Fragments Generated from Different Top-Down Mass Spectrometry Fragmentation Methods Extend Protein Sequence Coverage | Journal of the American Society for Mass Spectrometry

Build your own mass spectrometry analysis pipeline in Python using matchms — part I | by Florian Huber | Netherlands eScience Center

Pyteomics 4.0: Five Years of Development of a Python Proteomics Framework | Journal of Proteome Research

PypKa: A Flexible Python Module for Poisson–Boltzmann-Based pKa Calculations | Journal of Chemical Information and Modeling